This a brief guide to using R in collaborative, social ways. R is a powerful open-source programming language for data analysis, statistics, and visualization, but much of its power derives from a large, engaged community of users. This is an introduction to tools for engaging the community to improve your R code and collaborate with others.
(Am I missing anything? Let me know in the comments and I’ll update this guide.)
Topics
- Asking questions via e-mail, listservs and bulletin boards
- Sharing R scripts with
gist.github.com - Sharing reports of code and results with
knitr - Collaborating and version control
- Package creation and sharing
- RStudio
- Interactive R with Shiny
- Upcoming tools for real-time, interactive and collaborative programming
- Engaging with R Communities Online
Asking questions via e-mail, listservs and bulletin boards
One of the best ways to tap into the knowledge of the R community is to use the listservs and websites devoted to answering questions about R. There are a large number of these online forums. Some of the most popular are the R-help listserv, the progamming Q&A site Stack Overflow, and the statistics Q&A site Cross-Validated. (Be sure to look at the [R] tags on these two sites.) There are also specialty listservs like R-sig-ecology, or local forums like the Davis R Users’ Group.
These forums are great places to get help with your R questions. To get good answers, though its important to know how to ask good questions. Key to a good question is a minimal reproducible example (MRE). An MRE is a bit of code that, when copy-and-pasted from an e-mail into R, will reproduce the results or problem you are asking about. Here is a great guide to producing a reproducible example. A few important components:
- a minimal dataset, necessary to reproduce the error
- the minimal runnable code necessary to reproduce the error, which can be run on the given dataset.
- the necessary information on the used packages, R version and system it is run on.
- in case of random processes, a seed (set by set.seed()) for reproducibility
One useful trick for producing an MRE is the dput() command. dput takes any object in R and prints it in a form that can be copy-and-pasted. For instance, say you have a data frame like this:
## a b
## 1 1 6
## 2 2 7
## 3 3 8
## 4 4 9
## 5 5 10Running dput creates text that can be entered into R make an identical structure
## structure(list(a = 1:5, b = 6:10), .Names = c("a", "b"), row.names = c(NA,
## -5L), class = "data.frame")Now you can insert this data into your MRE by typing:
df <- structure(list(a = 1:5, b = 6:10), .Names = c("a", "b"),
row.names = c(NA, -5L), class = "data.frame")Your real data is probably considerably larger and more complex, and could be in the form of a data frame, list, or any other number of objects. Try running dput(mtcars) to see the results with a larger data set. It’s often shorter to use dput on cleaned-up, manipulated data than including a lot of data-manipulation steps in your MRE. If your dataset is large, simply use dput(head(mtcars)) to share only the first few rows. Here is an example of how using this method yields useful responses.
Similarly, it is easy to share more information about your set up using the sessionInfo() function. It returns information about your R version, your platform, base, and loaded packages, all of which are really helpful for troubleshooting purposes. Simply copy and paste the output from the function alongside your question.
Sharing R scripts with gist.github.com
Sometimes you want to share an R code that you’ve written. Creating a gist is a great way to do so. Gists are have several advantages over sharing over e-mail:
- You can share the code just by sending a URL
- Syntax highlighting makes the code easier to read
- You can update the code and it will remember previous versions
- The gist can be public or private
- People can comment on the gist and have a conversation
To post your code as a gist, go to http://gist.github.com, and paste your code, and put in a brief description. You’ll get an easily sharable web page like this:
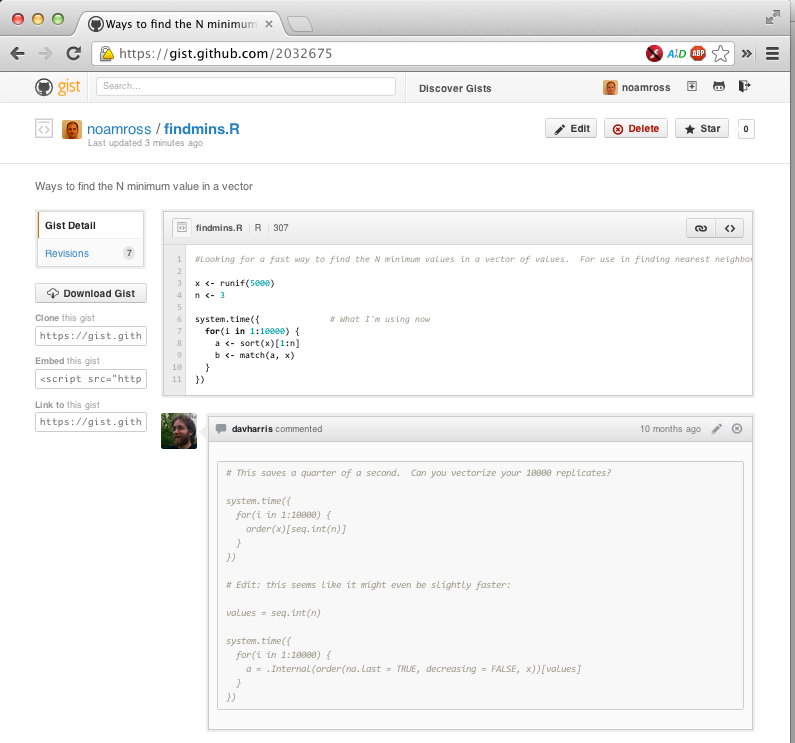
Signing up for the website is not neccessary, but it is needed if you want to revise your gist in the future.
A gist can include multiple files, so sometimes its useful for to include both an R script and your source data file in one gist.
Sharing reports of code and results with knitr
Often you want to share the results of analyses you perform in R with colleagues or the broader online community. A good way to do this is with a report - a document of text, code, and results (often graphs). knitr is an R package for making reports that can be printed or shared on the web.
knitr takes documents that are a mixture of text and code, extracts the code and runs it, and then inserts the code and results back in. This has several advantages:
- If you change your code, all you have to do is re-run knitr, rather than run the code, copy-and-paste results, and then do re-formatting
- Colleagues who read your report can see exactly the steps you took to reach your results, and reproduce them if they want.
knitr can work with a variety of different document types: HTML, LaTeX, etc., but most commonly people use markdown, which is a simple syntax designed for producing web pages. Here’s what a short markdown document using knitr looks like:
Title
=====
This is an example document. Here's a summary of some data:
```{r}
summary(cars)
```
Here is a plot of that data:
```{r}
plot(cars)
```If you save this file as example.Rmd (R-markdown), and run
library(knitr)
library(markdown)
knit("example.Rmd")
markdownToHTML("example.md")You get an HTML file called example.HTML. Open it in a browser and it looks like this:
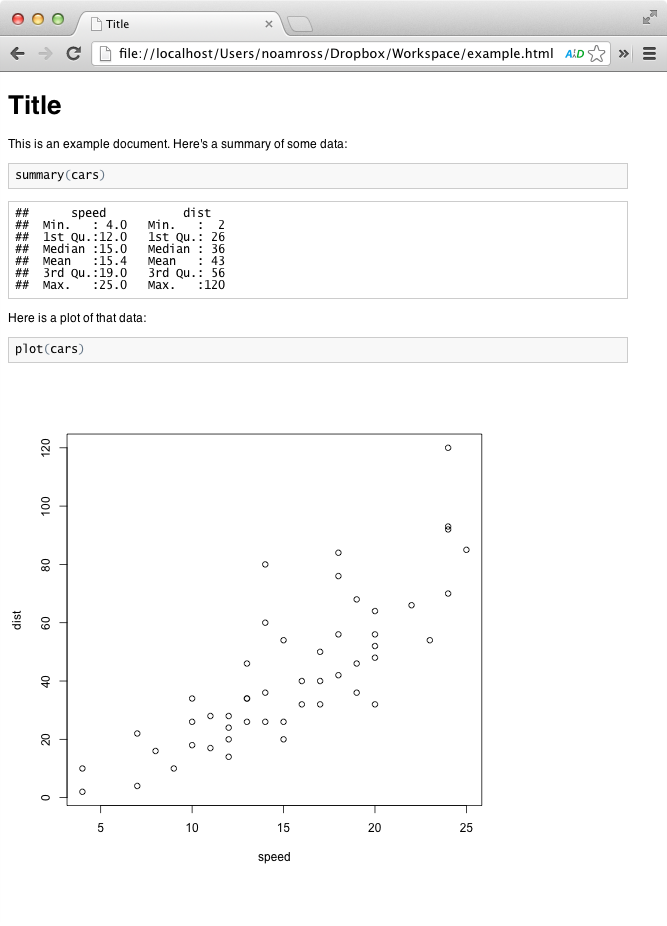
As you can see, your document has both your writing, code, and results, formatted in an easy-to-read way. You can e-mail these documents or post them online for easy sharing.
Collaborating and version control
If you are working with a team on a project that involves multiple R analyses and data sets, you’ll want a more robust system for collaboration that just e-mailing files around. Here are two options.
Dropbox
An easy way to share your R files and maintain version control is to use Dropbox. Dropbox is a service that syncs files across computers. If you and your collaborators share a Dropbox folder, changes will automatically stay up-to-date across all your computers. Importantly, Dropbox keeps previous versions of files and lets you revert to past versions, so you can go back if something in your code breaks:
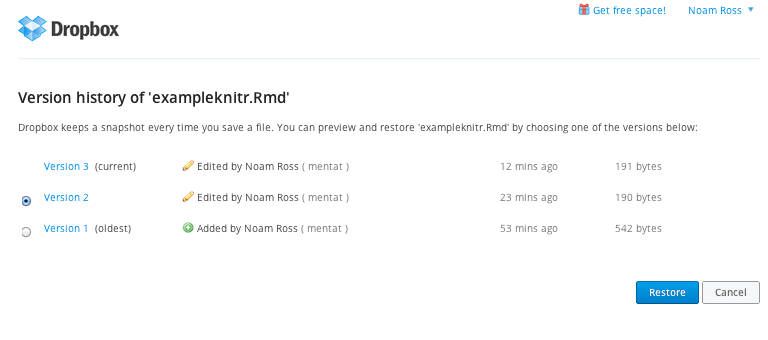
Git and Github
Git is a version-control system widely used by programmers. It’s much more powerful than Dropbox version control, with features that include
- Fine-scale control of what files and folders change with each update
- Log messages to remind you of what changes were important with each version
- Creating parallel versions (branches) of projects
- Comparing different versions of files and folders
- Merging changes made on different branches or by different users.
Git has a bit of a learning curve, but if you do a lot of programming work in R or other languages, it is well worth it. To get started, see the documentation here
Using Git alongside a web service like Github or Bitbucket lets you collaborate on projects in a very powerful way. Collaborators can work on code simultaneously, merge changes and resolve conflicts through the website. If you choose to make the project open-source, your code can be public and you can tap into the expertise of many other collaborators.
The R package devtools lets you download and run files directly from these websites.
Package creation and sharing
One of the great strengths of R is the collection of over 4,000 user created packages. You might want to create a package if you have developed a new method in R, or if you have a collection of helpful functions that would be useful to share. If you write a scientific publication using R for analysis, an accompanying package is a good way to make the data and methods accessible for other researchers to reproduce and build upon.
Hadley Wickham has written a great guide to developing R packages, which can be found here. It accompanies his package devtools which provides many useful tools for package development.
RStudio
If you want use the tools described above, it is helpful to use software that integrates them into your workflow. RStudio is a popular integrated development environment (IDE). An IDE makes working in R easier by putting the R console, a text editor, file browser, help files, graphics and many other tools together into a cohesive interface. RStudio also integrates many of the collaborative tools described above. It is available for Mac, Windows, and Linux.
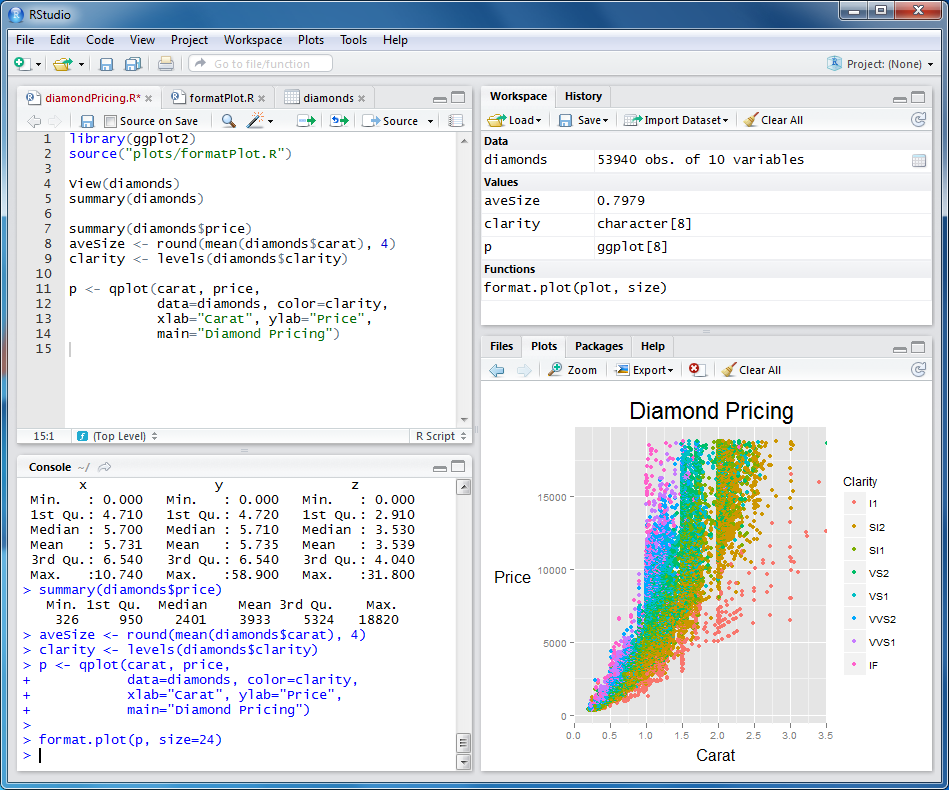
If you use Rstudio, knitr is automated for you. Just hit the “Knit HTML” button once you have written your R-markdown document, and it will generate the web page and show a preview:
An additional benefit of using knitr from Rstudio is that it will give you the option of automatically uploading the HTML document to their server at http://rpubs.com/ so you can share it with anyone. Just hit the “Publish” in the preview window:
Like gists, documents at http://rpubs.com/ can be updated and easily shared and accept comments.
Git is also built into the Rstudio interface. Saving a version of your software is as easy as clicking a button. This makes the git learning curve a little easier:
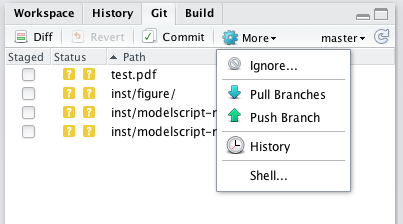
Finally, the latest version of Rstudio has package creation tools based on the devtools package, including tools for testing and documenting packages. Like its Git interface, RStudio’s package development tools make the process of package creation more intuitive:
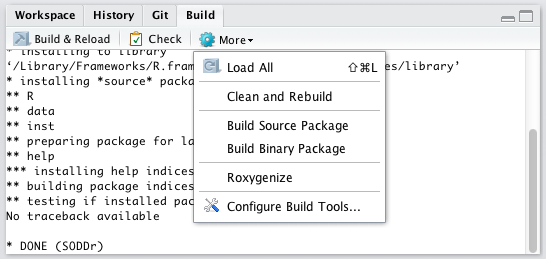
Interactive R with Shiny
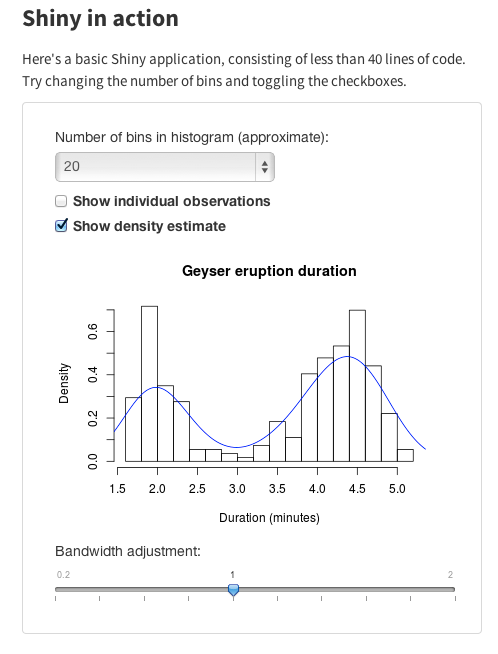
A relatively new and exciting way to use R to share data analysis is Shiny. Shiny is an R package that lets you create interactive web pages that let users explore your data and analysis. Here’s an example. Click on it to go to the interactive version:
Upcoming tools for real-time, interactive and collaborative programming
In the next year or so we will likely see tools for live, interactive coding that allow you to collaborate in real time on R scripts the way Google Docs allows such collaboration with documents. https://www.stypi.com is one such tool, though it doesn’t have R-specific options yet. Yihui Xie, the creator of knitr has created an interactive notebook based on Shiny and knitr which runs your knitr/R code on the web, but it is just a proof-of-concept. Look out for new developments!
Engaging with R Communities Online
If you are looking for peers and collaborators in your work with R, there are a lot of places online to do so.
Listservs:In addition the listservs mentioned above, there are many specialty listservs for specific platforms and applications, and local user groups. Many packages have listservs associated with them, too, where users can ask question and get information about the latest updates. If you are learning how to use a new package it’s helpful to sign up for these.
Blogs: R-bloggers is a great website that aggregates many blogs by people who use R. Blogs range from people writing about R (e.g., tutorials), to people blogging with R (e.g., knitr documents of their latest analyses). Following the site feed will help you discover other people doing work similar to yours.
Twitter: If you’re on twitter, the #rstats hashtag is commonly used to discuss R, and you can often get answers to short questions very quickly. Many of developers of R packages and software are on twitter, so you can get information straight from the source.
Code Hosting Sites: Many R package developers host their projects on websites like Github and R-forge. These sites have mechanisms for users to report bugs, make feature requests, and often find more information than is available in the documentation of a specific package. You can often find the site used by a developer by looking at it’s entry on CRAN or the package revewing site crantastic.